What is a gene family in PhyloGenes?
Gene families in PhyloGenes are pruned versions of PANTHER gene families (pantherdb.org, Mi2019). They contain only genes from selected plant genomes and 10 non-plant model organisms (phylogenes.org). Genes from other genomes in the PANTHER build have been removed (pruned) from the PANTHER gene families and gene trees.
A PANTHER gene family contains genes that are related to each other by descent from a common ancestor, as established by statistical sequence similarity, and whose protein sequences can be aligned reliably into a multiple sequence alignment. The UniProt Reference Proteomes used in PANTHER family construction contain one representative protein sequence per gene. A gene family is represented as a phylogenetic gene tree that shows how the family evolved by the processes of speciation, gene duplication, and horizontal transfer.
How is the boundary of a gene family defined?
In PANTHER, gene families are defined as clusters of related protein sequences (each protein sequence represents a distinct gene) for which a good multiple sequence alignment can be made (Mi2013, Mi2016). The basic requirements for a family are: (1) the family contains at least five sequences and includes more than one organism, and (2) the family has a sequence alignment of adequate quality to support phylogenetic inference. An alignment must have at least 30 sites aligned across 75% or more of the family members, and the derived Hidden Markov Model (HMM) must be able to recognize, with statistical significance, the sequences used to train it.
What is the procedure you used in building a gene family and constructing its gene tree?
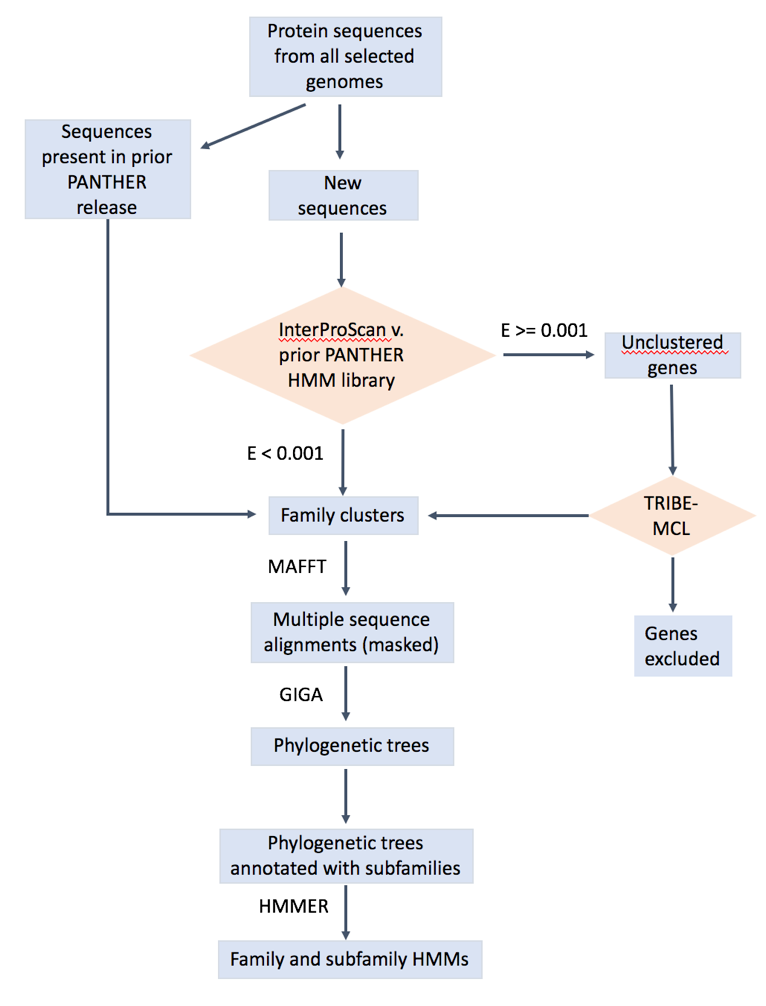
The overall workflow is shown below. The details can be found in Mi2013, Mi2016, Mi2017 .
How is a PANTHER gene tree constructed?
A PANTHER gene tree is composed of orthologous subtrees (containing protein sequences related by speciation events), joined together by gene duplication (duplication within the same genome) or horizontal transfer (insertion from another genome). PANTHER trees were constructed by using the GIGA algorithm (Thomas2010). The algorithm builds a tree from leaf nodes to the root, using a pairwise sequence distance matrix, a known species tree (based on NCBI taxonomy), and a set of rules to establish the tree topology. Sequence distance is calculated as the fraction of sequence differences between two sequences at selected homologous sites. The homologous sites were selected from multiple sequence alignment of all genes in the family. GIGA iteratively joins together subtrees of sequences, beginning with the two sequences that are closest according to the pairwise sequence distance matrix. The topology of the joined subtree after each iteration is not simply an agglomeration of the constituent subtrees. Rules are used to "rearrange" the joined subtree at each iteration. For example, if a subtree contains only speciation events, the topology is determined by the known species tree. Copying events such as duplication or horizontal transfer are placed within a tree with the most parsimonious solution to minimize gene deletions. The full description of the GIGA algorithm can be found in this paper (Thomas2010).
How is branch length calculated?
First, the ancestral sequence is inferred for each non-leaf node using a local, parsimony-like algorithm that reconstructs each node using only its descendants and closest outgroup. If over half of the descendant nodes align the same amino acid at a given site, it is inferred to be the most likely ancestral amino acid. If the descendants disagree, and the outgroup agrees with one of them, the outgroup amino acid is inferred to be the most likely ancestral amino acid. Otherwise, the ancestral amino acid is considered to be unknown ('X'). Next, the branch length between a parent node and a child node was calculated as the fraction of sequence differences between them. The Jukes-Cantor correction is applied to this value. More details can be found here (Thomas2010).
Why aren’t there any bootstrap or other support values for the tree topology?
Although the GIGA algorithm builds trees using a distance matrix, the topology of a tree is less influenced by sequences when compared to other distance or character-based algorithms such as neighbor joining or maximum likelihood. The reason is that GIGA imposes strict rules in determining tree topology. It uses a pre-established species tree in sorting out speciation nodes. GIGA uses maximum parsimony in placing duplication nodes. Therefore, the topology of a tree is very robust. For example, the rule of using species tree is illustrated below:
What is a subfamily?
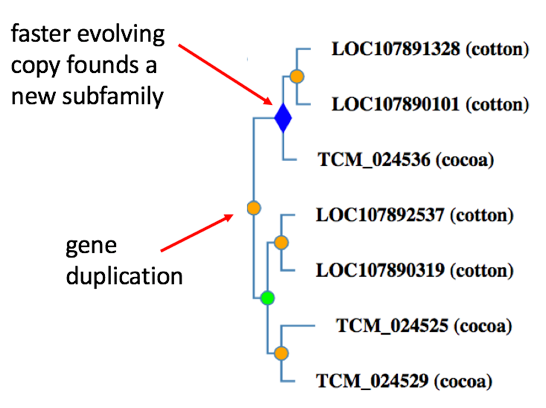
Subfamilies within each family are groups of genes that share a particularly high degree of protein sequence similarity due to limited divergence from their common ancestor (Mi2016). Subfamilies are, in general, closely-related orthologs. In the PANTHER tree building process, a new subfamily is created within a family after every gene duplication event, or horizontal transfer event. After horizontal transfer, the transferred copy becomes the founder of a new subfamily; the vertically inherited copy remains in the original subfamily. After gene duplication, the copy that changes faster in sequence immediately following the duplication becomes the founder of a new subfamily; the slower-evolving copy remains in the same subfamily. There are two exceptions to this rule: (1) because of the high frequency of gene duplication prior to the vertebrate common ancestor, each vertebrate copy following a gene duplication event founds a new subfamily, and (2) duplicated genes do not found a subfamily if they did not lead to orthologs in at least two extant species.
How are gene families named? Why are some families not named?
Biologically meaningful family names are assigned by biologist curators (Thomas2003). The curator either assigns a family a more general functional name that applies to all genes in the family (e.g., NUCLEAR HORMONE RECEPTOR) or finds the largest subfamily name (Y) and names the family Y-RELATED. If in the latter case the largest subfamily is “unnamed” then the family is not named.
How are subfamilies named? Why are some subfamilies not named?
The name of a subfamily is transferred from the representative member of the subfamily (Mi2016). If a subfamily contains an annotated SwissProt entry from any of the 12 model organisms (human, mouse, rat, chicken, zebrafish, fruit fly, C. elegans, budding yeast, fission yeast, D. discoideum, Arabidopsis, and E. coli.), then the curated ‘protein name’ is used to name the subfamily. If a subfamily does not contain any SwissProt entries from the model organisms, but contains SwissProt entries from other organisms, the most common SwissProt protein name is used as the subfamily name. If a subfamily does not contain any members from the SwissProt database, a protein name from the TrEMBL entry is automatically selected as the subfamily name. If no name can be found, the subfamily is labeled with ‘unnamed’.
What is horizontal gene transfer? How is it detected in PANTHER gene trees?
Horizontal transfer is the movement of genetic material between organisms other than by the default "vertical" transmission from parent to offspring. Horizontal transfer is common in bacteria. In Eukaryotes, events such as the engulfment of the mitochondrion by the proto-eukaryotic cell will be represented at the level of gene family trees as horizontal transfer from a proteobacterial ancestor to the eukaryotic common ancestor. At each step in PANTHER gene tree building the GIGA algorithm considers the number of gene deletions that would be implied by a history of vertical inheritance, and if that number is too large, a horizontal transfer event is considered to be the more likely interpretation (Mi2016).
Are polyploid organisms represented in gene families?
Several polyploid organisms are included in PhyloGenes. Just like genes from a single genome in a diploid, genes from N (N>1) genomes of a polypoid are treated as individual genes. For example, the bread wheat is a hexaploid, which has three genomes A, B and D. GeneX is present in all three genomes therefore there are three copies of GeneX in bread wheat, GeneX-A, GeneX-B and GeneX-D. Given high sequence similarities shared among the three genes, GeneX-A, GeneX-B and GeneX-D are very likely to be found in the same gene tree shown as descendants of a gene duplication event.
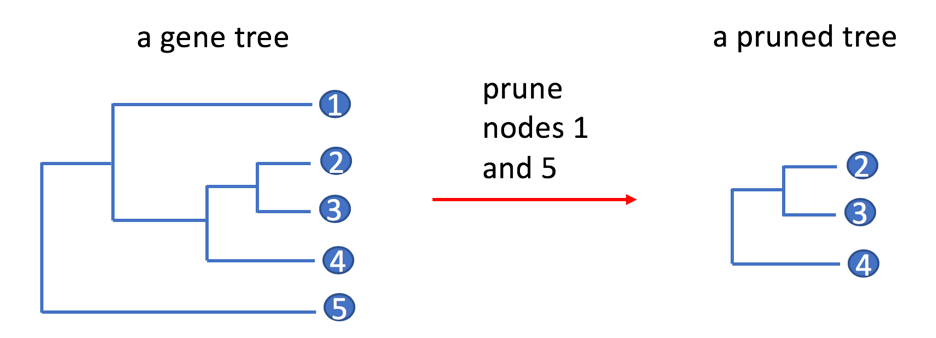
How does tree pruning work? Does it change the topology of a tree?
Gene families in PhyloGenes are pruned versions of PANTHER gene families. They contain only genes from selected plant genomes and non-plant model organisms. Genes from other PANTHER genomes were removed (pruned) from the PANTHER gene families and phylogenetic trees. The pruning process does not alter the tree topology of a gene family. Given a PANTHER gene tree, the process simply removes a leaf node (gene) if the gene is NOT from any of the selected organisms.
My gene is not found in PhyloGenes. Why?
There are two possibilities. It could be that the organism of your gene is not included in the PANTHER pipeline. The list of genomes that are included PANTHER and PhyloGenes can be found here. Alternatively, even if the organism is included in PANTHER and PhyloGenes, not all genes of the genome are part of a gene family. Some genes don’t meet the criteria of being part of a gene family. Statistics covering the current release can be found here.


Add Comment